Protein expression systems are used to produce proteins for use in research, development, and clinical studies. Boster Bio’s protein expression services use these protein expression systems to produce proteins from genes and other DNA sequences. These services include the creation of custom protein expression vectors and the purification of the resulting protein.
Protein expression systems are biological systems used to produce recombinant proteins. They can be prokaryotic, eukaryotic, or both. These systems can be used to study and modify the function of proteins, although they have other uses as well. Boster offers custom recombinant protein production services and bacterial protein expression services.
What is Protein Expression?
https://encrypted-tbn0.gstatic.com/images?q=tbn:ANd9GcSPKVB7UuehuaHLpulTLjwGulBsyIwGeTSqGg&usqp=CAU
Protein expression is the process of making a protein from a gene. It involves copying the gene into messenger RNA (mRNA) molecules, which are then translated into protein molecules. In order to make this happen, you’ll need a protein expression system.
What is a protein expression system?
A protein expression system is a method for producing proteins from DNA molecules, or genes. The process of protein expression involves the use of a vector (plasmid) to carry DNA from one cell to another. The plasmid acts as an intermediate in the transfer of genetic information from one cell to another, including artificial DNA. Proteins are produced by expressing specific genes that code for proteins within living cells, which can be grown in large quantities in Petri dishes or through fermentation processes.
The ability to produce proteins quickly and efficiently has become essential for many industries because it allows them to make products more cheaply than they could otherwise. It also helps scientists study new processes related to biological systems without having to wait years for them to occur naturally. The most common expression system is E. coli, which works for many protein targets but not all; other common systems include yeast, mammalian cells, and plants.
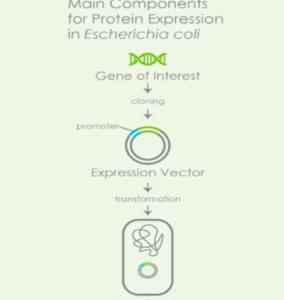
How are recombinant proteins expressed in E. coli?
The E. coli cell produces the protein using its own machinery, so you don’t need to worry about producing it yourself. The DNA sequence encoding the protein is inserted into a plasmid, which can be easily purified away from contaminants and then introduced into the E. coli strain by transformation or conjugation.
The E. coli strain is grown in a culture medium that contains all the necessary nutrients for growth except oxygen; if glucose is used as the carbon source, fermentation can occur in which glucose is converted to ethanol and acid products by alcohol dehydrogenase enzymes produced by the bacterial cells themselves.
The expression of recombinant proteins takes place in the cytoplasm of E. coli, where they are secreted into the culture medium after being synthesised on ribosomes attached to membrane-bound ribosomes within vesicles known as “assembly sites” (or sometimes “vesicular bodies”).
These vesicular bodies fuse with late endosomes containing acidic pH conditions which cause denaturation of hydrophobic peptide bonds needed for correct folding. This process also exposes hydrophilic regions that allow them access out of these vesicles through specialised pores called porins located on their surface.
There are multiple ways to express proteins in E. coli, but the most commonly used method is called the “strep” system. This involves growing the bacteria in a culture medium containing glucose and other nutrients, adding a gene of interest to the cell’s chromosome, then allowing it to replicate until enough copies of that gene have been made. The protein itself is synthesised by ribosomes within cells and secreted into the surrounding environment through one of two channels.
Once outside of a cell, recombinant proteins must be purified from their natural milieu using an affinity column; this step ensures that only recombinant molecules will be isolated from solution during downstream processing steps such as chromatography or electrophoresis.
The most common expression system is E. coli
- coli is a prokaryotic organism and the most common expression system used in biotechnology. It’s also a fast-growing bacteria that’s easily manipulated by scientists, making it ideal for use as an expression system. The ease of manipulation makes E. coli an excellent choice for both basic research and large-scale production of proteins or other biomolecules (such as antibodies).
- coli is often referred to as being gram negative because its outer membrane contains lipopolysaccharides (LPS), which are negatively charged polysaccharides that are responsible for the cell’s impermeability to many substances including certain antibiotics. Because of this, many antibiotics can’t penetrate into E. coli cells; however some have been developed specifically for use with E. coli like streptomycin, which has been used since 1952 to treat tuberculosis infections caused by M tuberculosis (caused by mycobacterium intracellularis). Other common protein expression systems include: yeast expression system, insect expression system, mammalian cell expression system
The decision regarding which system to use depends on the properties of your protein of interest. Se determining Factors include:
Protein size
If you have a relatively small protein, such as a monomeric protein or a polypeptide, then a transient expression system may be suitable for your work. If you have a large multimeric protein, like an antibody or enzyme, then prokaryotic vectors may be more appropriate.
Protein Structure
If the native conformation of your protein is important in its function (i.e., it folds into some unique shape), then this factor should also be considered when choosing an expression system since most mammalian cells lack the machinery needed to fold proteins correctly and will likely produce mis-folded forms if expressed at high levels.
Conclusion
As you can see, there are many factors to consider when choosing an expression system for the protein you’re working with. While E. coli is the most common bacterial host, this may not be appropriate for your research needs. Consider your options carefully and select a system that best fits your project’s requirements.
